I1 and/or I2 FASTQ files are optional.įor more information on FASTQ format requirements, please see Specifying FASTQ files. Note that only R1 and R2 FASTQ files are required for Cell Ranger. compatible file name: SRR9291388_S1_R1_ (for Cell Ranger v4.0 and later)Ĭhanging the file names will allow Cell Ranger (version >=2.1.1) to accept these data as input.compatible file name: SRR9291388_S1_L001_R1_ Sequence Read Archive (SRA) data, available through multiple cloud providers and NCBI servers, is the largest publicly available repository of high throughput sequencing data.incompatible file name: SRR9291388_1.fastq.gz.NOTE: Cell Ranger v4.0 and later will accept file names without lane number. The number of FASTQ files we retrieved is consistent with what is reported in the metadata of SRR9291388: This run has 3 reads per spot.īased on the read length and the attributes listed in the metadata, we know that:Ĭell Ranger requires FASTQ file names to follow the bcl2fastq file naming convention. The output would be three FASTQ files: SRR9291388_1.fastq.gz For example, if we try to retrieve FASTQ files from SRR9291388: fastq-dump -split-files -gzip SRR9291388 Sometimes, the authors may upload 3 or 4 reads for a sample by including index reads. Download and convert SRA files to FASTQ files using the NCBI’s SRA toolkit. The number of FASTQ files we retrieved is consistent with what is reported in the metadata of SRR6334436: This run has 2 reads per spot.īased on the read length, we can infer that SRR6334436_1.fastq is Read 1 and SRR6334436_2.fastq is Read 2 (see Sequencing Recommendation for 3' Gene Expression Assays) The output would be two FASTQ files: SRR6334436_1.fastq.gz 
The command may look like this: fastq-dump -split-files -gzip SRR6334436 The following fix has been tested on Chromium v2 and v3 chemistry.įirst, use the NCBI fastq-dump utility with the -split-files argument to retrieve the FASTQ files. The prefetch - tool can be invoked multiple times if the download did not succeed. The prefetch tool downloads all necessary files to your computer. The combination of prefetch + fasterq-dump is the fastest way to extract FASTQ-files from SRA-accessions. Before downloading SRA data, first identify the platform and version of the chemistry used to generate the data. How to use prefetch and fasterq-dump to extract FASTQ-files from SRA run accessions. Using these data in Cell Ranger requires some pre-processing. The primary source of these publicly available datasets in the United States is the Sequence Read Archive (SRA) maintained by NCBI. 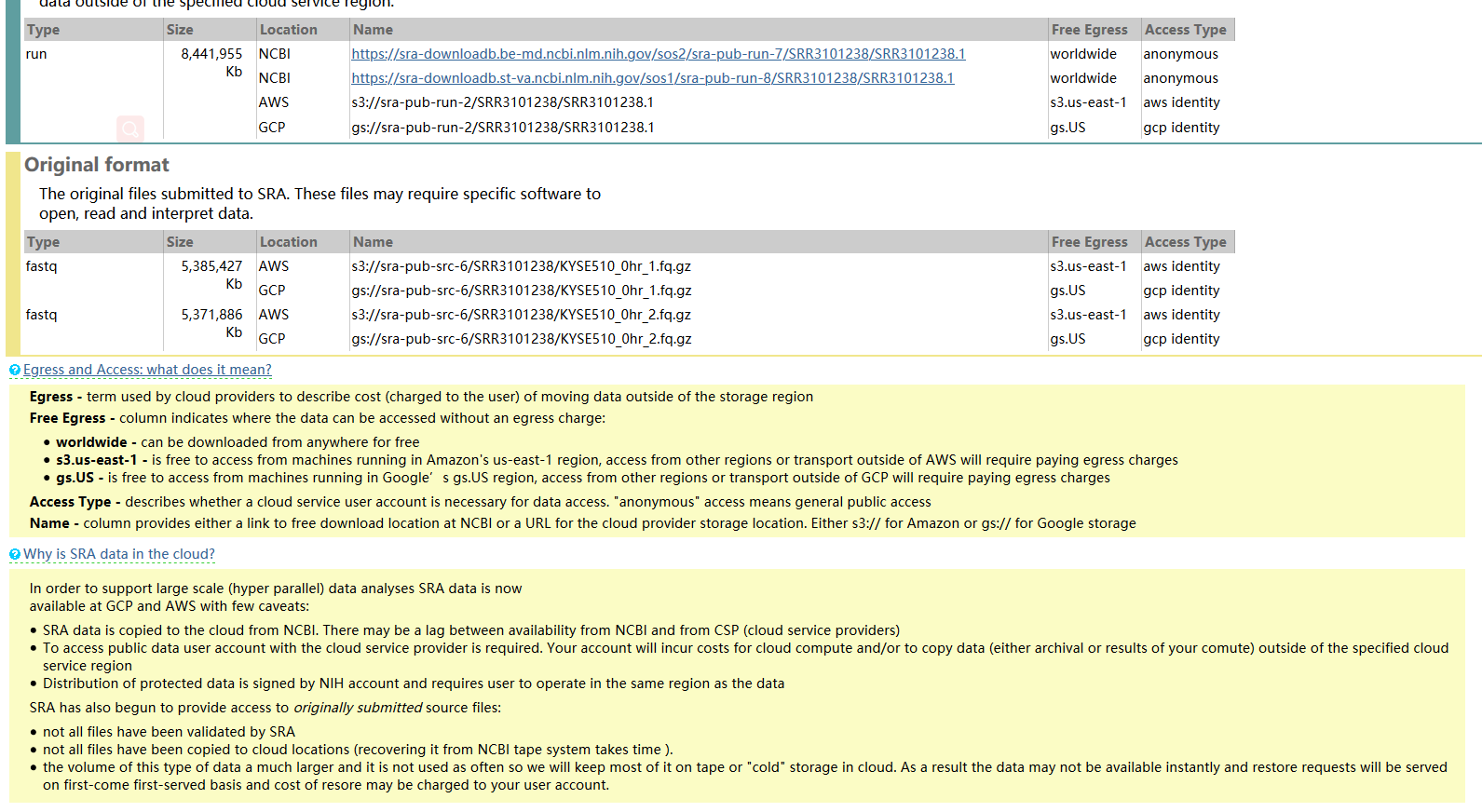
Question: How do I prepare Sequence Read Archive (SRA) data from NCBI for Cell Ranger?Īnswer: One of the beauties of open source data in the sequencing age is the ability to reanalyze data generated by other researchers.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
